Tidyverse and Some SQLite
Overview
Modern object classes and methods for handling data.frame like structures are provided by the dplyr (tidyr) and data.table packages. A related example is Bioconductor’s DataTable object class (“Learn the tidyverse,” n.d.). This tutorial provides a short introduction to the usage and functionalities of the dplyr and related packages.
Why learn both base R and tidyverse?
Most genomics and Bioconductor package documentation is written in base R, which is why GEN242 introduces base R syntax first and focuses on it throughout the course. The tidyverse (centered on dplyr, tidyr, and ggplot2) offers an alternative syntax that many find more readable for interactive data exploration, and you will encounter it frequently in online tutorials, Stack Overflow answers, and newer analysis workflows. This tutorial introduces the key dplyr functions side-by-side with their base R equivalents so you can read and write both styles fluently.
A rough comparison of the three main approaches:
| Feature | Base R | dplyr / tidyverse | data.table |
|---|---|---|---|
| Learning curve | Moderate | Low–moderate | Steep |
| Verbosity | More verbose | Concise, readable | Very concise |
| Performance (large data) | Moderate | Moderate | Excellent |
| Bioconductor docs | ✓ Primary style | Occasionally | Rarely |
| Pipe-friendly | With \|> (R ≥ 4.1) |
Core design | Partial |
The data.table package is not covered in depth here, but its fread() function is introduced for fast file import, as it is widely used even in tidyverse workflows.
Installation
The dplyr (tidyr) environment has evolved into an ecosystem of packages. To simplify package management, one can install and load the entire collection via the tidyverse package. For more details on tidyverse see here.
Construct a tibble (tibble)
Reading and writing tabular files
While the base R read/write utilities can be used for data.frames, best time performance with the least amount of typing is achieved with the export/import functions from the readr package. For very large files the fread function from the data.table package achieves the best time performance.
Import with readr
Import functions provided by readr include:
read_csv(): comma separated (CSV) filesread_tsv(): tab separated filesread_delim(): general delimited filesread_fwf(): fixed width filesread_table(): tabular files where columns are separated by white-space.read_log(): web log files
Create a sample tab delimited file for import
Import with read_tsv
To import Google Sheets directly into R, see here.
Fast table import with fread
The fread function from the data.table package provides the best time performance for reading large tabular files into R.
library(data.table)
iris_df <- as_tibble(fread("iris.txt")) # Import with fread and conversion to tibble
iris_dfNote: to ignore lines starting with comment signs, one can pass on to fread a shell command for preprocessing the file. The following example illustrates this option.
Export with readr
Export functions provided by readr include
write_delim(): general delimited fileswrite_csv(): comma separated (CSV) fileswrite_excel_csv(): excel style CSV fileswrite_tsv(): tab separated files
For instance, the write_tsv function writes a data.frame or tibble to a tab delimited file with much nicer default settings than the base R write.table function.
Column and row binds
The equivalents to base R’s rbind and cbind are bind_rows and bind_cols, respectively.
Extract column as vector
The subsetting operators [[ and $can be used to extract from a tibble single columns as vector.
Important dplyr functions
filter()andslice()arrange()select()andrename()distinct()mutate()andtransmute()summarise()sample_n()andsample_frac()
Slice and filter functions
Filter function
Base R code equivalent
Including boolean operators
Subset rows by position
dplyr approach
Base R code equivalent
Subset rows by names
Since tibbles do not contain row names, row wise subsetting via the [,] operator cannot be used. However, the corresponding behavior can be achieved by passing to select a row position index obtained by basic R intersect utilities such as match.
Create a suitable test tibble
df1 <- bind_cols(tibble(ids1=paste0("g", 1:10)), as_tibble(matrix(1:40, 10, 4, dimnames=list(1:10, paste0("CA", 1:4)))))
df1dplyr approach
Base R equivalent
Sorting with arrange
Row-wise ordering based on specific columns
dplyr approach
For ordering descendingly use desc() function
Base R code equivalent
Select columns with select
Select specific columns
Select range of columns by name
Drop specific columns (here range)
Change column order with relocate
dplyr approach
For details and examples see ?relocate
Base R code approach
Renaming columns with rename
dplyr approach
Base R code approach
Obtain unique rows with distinct
dplyr approach
Base R code approach
Add columns
mutate
The mutate function allows to append columns to existing ones.
transmute
The transmute function does the same as mutate but drops existing columns
bind_cols
The bind_cols function is the equivalent of cbind in base R. To add rows, use the corresponding bind_rows function.
Summarize data
Summary calculation on single column
Summary calculation on many columns
Summarize by grouping column
Aggregate summaries
Note: group_by does the looping for the user similar to aggregate or tapply.
Merging tibbles
The dplyr package provides several join functions for merging tibbles by a common key column similar to the merge function in base R. These *_join functions include:
inner_join(): returns join only for rows matching among bothtibblesfull_join(): returns join for all (matching and non-matching) rows of twotibblesleft_join(): returns join for all rows in firsttibbleright_join(): returns join for all rows in secondtibbleanti_join(): returns for firsttibbleonly those rows that have no match in the second one
Sample tibbles to illustrate *.join functions.
df1 <- bind_cols(tibble(ids1=paste0("g", 1:10)), as_tibble(matrix(1:40, 10, 4, dimnames=list(1:10, paste0("CA", 1:4)))))
df1df2 <- bind_cols(tibble(ids2=paste0("g", c(2,5,11,12))), as_tibble(matrix(1:16, 4, 4, dimnames=list(1:4, paste0("CB", 1:4)))))
df2Inner join
Note: dplyr 1.1.0+ introduced join_by(ids1 == ids2) as an alternative to the by=c("ids1"="ids2") syntax shown here. Both forms work identically; the older named-vector form is used throughout this tutorial as it remains the most common style in existing genomics code.
Left join
Right join
Full join
Anti join
For additional join options users want to consult the *_join help pages.
Chaining
To simplify chaining of several operations, dplyr uses the %>% operator from magrittr, where x %>% f(y) turns into f(x, y). This way one can pipe together multiple operations by writing them from left-to-right or top-to-bottom. This makes for easy to type and readable code. Since R-4.1.0, a native pipe |> operator is available that works largely the same as %>%. The native pipe requires no package and is the preferred style in modern base R code. Both forms will appear in online examples, so it is helpful to be comfortable with either:
# magrittr pipe (tidyverse style)
iris_df %>% filter(Species == "setosa") %>% nrow()
# Native pipe (base R >= 4.1, no package needed)
iris_df |> filter(Species == "setosa") |> nrow()The main practical difference is that |> does not support the . placeholder that %>% allows. For the examples in this tutorial the two are interchangeable.
Example 1
Series of data manipulations and export
read_tsv("iris.txt") %>% # Import with read_tsv from readr package
as_tibble() %>% # Declare to use tibble
select(Sepal.Length:Species) %>% # Select columns
filter(Species=="setosa") %>% # Filter rows by some value
arrange(Sepal.Length) %>% # Sort by some column
mutate(Subtract=Petal.Length - Petal.Width) # Calculate and appendExample 2
Series of summary calculations for grouped data (group_by)
Example 3
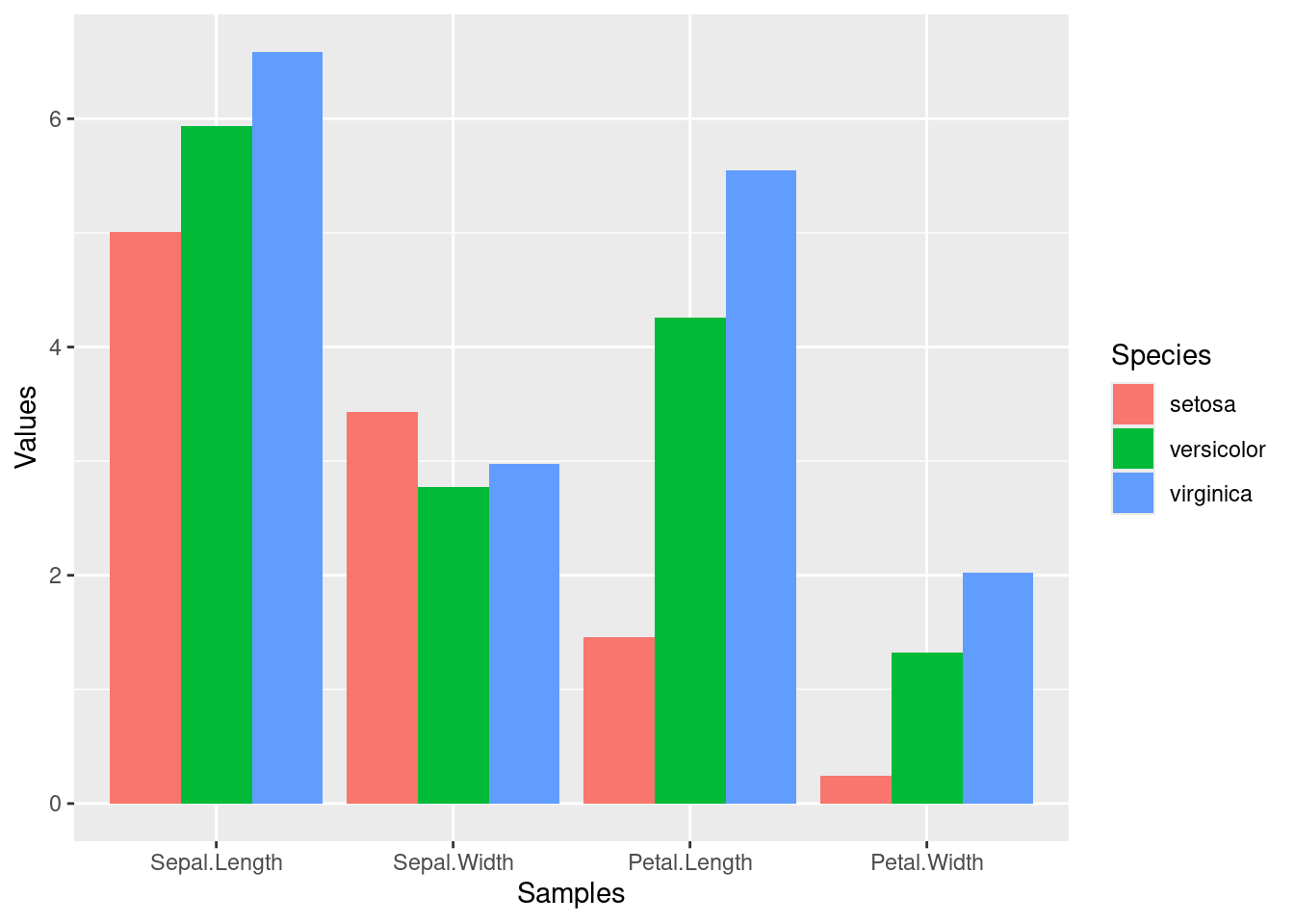
Combining dplyr chaining with ggplot
SQLite Databases
SQLite is a lightweight relational database solution. The RSQLite package provides an easy to use interface to create, manage and query SQLite databases directly from R. Basic instructions for using SQLite from the command-line are available here. A short introduction to RSQLite is available here.
Why SQLite matters in genomics
Many Bioconductor annotation packages — such as org.Hs.eg.db, TxDb.*, and BSgenome companions — store their data internally as SQLite databases. The AnnotationDbi package queries these databases on your behalf, but understanding how SQLite works helps you write more efficient annotation queries, troubleshoot unexpected results, and build your own lightweight databases for project-specific metadata (e.g. sample sheets, gene sets, experimental conditions). The examples below use a small iris-derived database to illustrate the core operations.
Loading data into SQLite databases
The following loads two data.frames derived from the iris data set (here mydf1 and mydf2) into an SQLite database (here test.db).
library(RSQLite)
unlink("test.db") # Delete any existing test.db
mydb <- dbConnect(SQLite(), "test.db") # Creates database file test.db
mydf1 <- data.frame(ids=paste0("id", seq_along(iris[,1])), iris)
mydf2 <- mydf1[sample(seq_along(mydf1[,1]), 10),]
dbWriteTable(mydb, "mydf1", mydf1)
dbWriteTable(mydb, "mydf2", mydf2)List names of tables in database
Import table into data.frame
Query database
Join tables
The two tables can be joined on the shared ids column as follows.
Disconnect from database
Always close the database connection when finished to release file locks and flush any pending writes.
Exercises
The following exercises reinforce the dplyr and SQLite concepts covered in this tutorial. For each dplyr task, try writing the solution in both dplyr and base R syntax.
Exercise 1 — Filtering and sorting
Using the iris dataset, select all rows where Petal.Width > 1.5 and Species is "virginica" or "versicolor". Sort the result by Species (ascending) and then by Petal.Width (descending). Show only the Species, Petal.Length, and Petal.Width columns.
Exercise 2 — Grouped summaries
For each Species in iris, compute the mean and standard deviation of Sepal.Length and Sepal.Width. Store the result as a tibble and inspect it.
Exercise 3 — Mutate and pipe chaining
Using a pipe chain, add a column Petal.Ratio (= Petal.Length / Petal.Width) to iris, filter to rows where Petal.Ratio > 3, and then return only the top 10 rows sorted by Petal.Ratio (descending).
Exercise 4 — Joining tibbles
Create a small annotation tibble:
annot <- tibble(
Species = c("setosa", "versicolor", "virginica"),
CommonName = c("Bristle-pointed iris", "Blue flag iris", "Virginia iris")
)Join annot to the iris tibble so that every row in iris gains a CommonName column. Which join type is appropriate here, and why?
Exercise 5 — SQLite
Re-open (or re-create) the test.db database from the tutorial. Write an SQL query that returns all rows from mydf1 where Species is "virginica" and Sepal.Length > 6.5. Then write the result back to the database as a new table called virginica_large. Verify the new table exists with dbListTables(), and close the connection when done.
Session Info
R version 4.5.3 (2026-03-11)
Platform: x86_64-pc-linux-gnu
Running under: Debian GNU/Linux 12 (bookworm)
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.11.0
LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.11.0 LAPACK version 3.11.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8 LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8 LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: America/Los_Angeles
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] RSQLite_2.4.6 data.table_1.18.2.1 lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.0 purrr_1.2.1 readr_2.1.6 tidyr_1.3.2 tibble_3.3.1
[11] tidyverse_2.0.0 ggplot2_4.0.2 limma_3.66.0
loaded via a namespace (and not attached):
[1] generics_0.1.4 stringi_1.8.7 hms_1.1.4 digest_0.6.39 magrittr_2.0.4 evaluate_1.0.5 grid_4.5.3 timechange_0.4.0 RColorBrewer_1.1-3 blob_1.3.0 fastmap_1.2.0
[12] plyr_1.8.9 jsonlite_2.0.0 DBI_1.2.3 scales_1.4.0 codetools_0.2-20 cli_3.6.5 rlang_1.1.7 crayon_1.5.3 bit64_4.6.0-1 cachem_1.1.0 withr_3.0.2
[23] yaml_2.3.12 otel_0.2.0 tools_4.5.3 parallel_4.5.3 reshape2_1.4.5 tzdb_0.5.0 memoise_2.0.1 vctrs_0.7.1 R6_2.6.1 lifecycle_1.0.5 htmlwidgets_1.6.4
[34] bit_4.6.0 vroom_1.7.0 pkgconfig_2.0.3 pillar_1.11.1 gtable_0.3.6 Rcpp_1.1.1 glue_1.8.0 statmod_1.5.1 xfun_0.56 tidyselect_1.2.1 knitr_1.51
[45] dichromat_2.0-0.1 farver_2.1.2 htmltools_0.5.9 labeling_0.4.3 rmarkdown_2.30 compiler_4.5.3 S7_0.2.1