library(gplots)
y <- matrix(rnorm(500), 100, 5, dimnames=list(paste("g", 1:100, sep=""), paste("t", 1:5, sep="")))
heatmap.2(y) # Shortcut to final result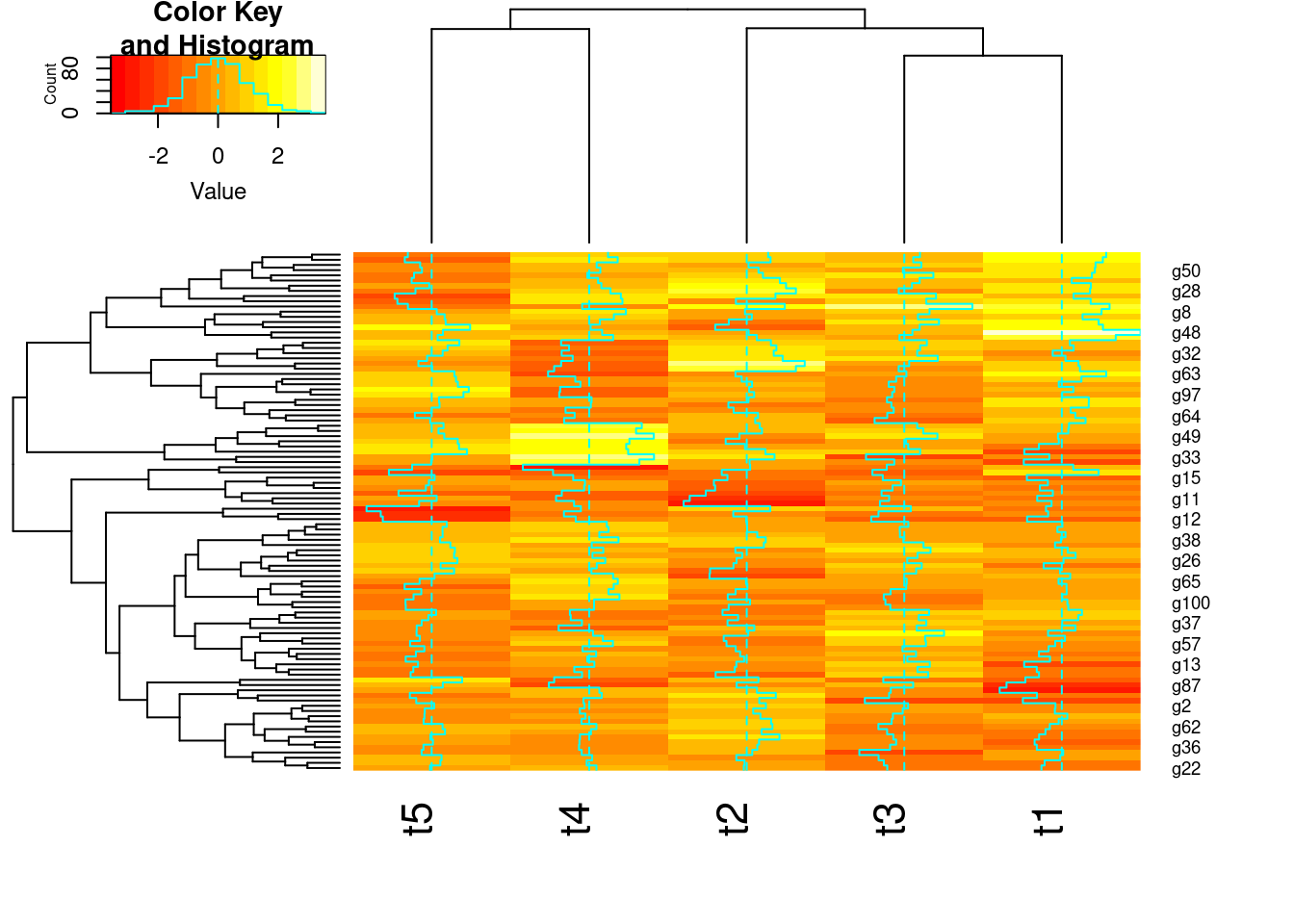
First/last name (first.last@ucr.edu)
May 21, 2026
Choice depends on data set!
scale() function
List of most common ones!
There are many more distance measures
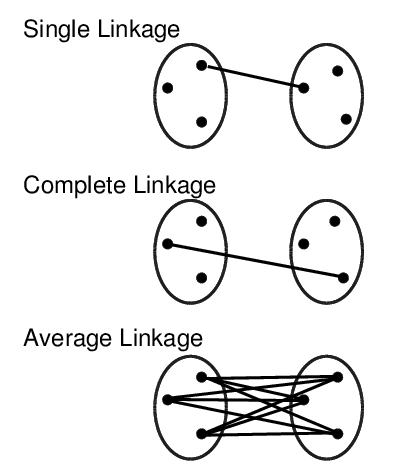
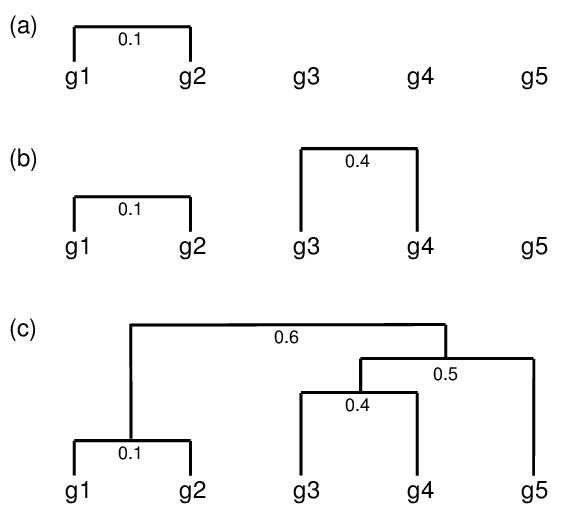
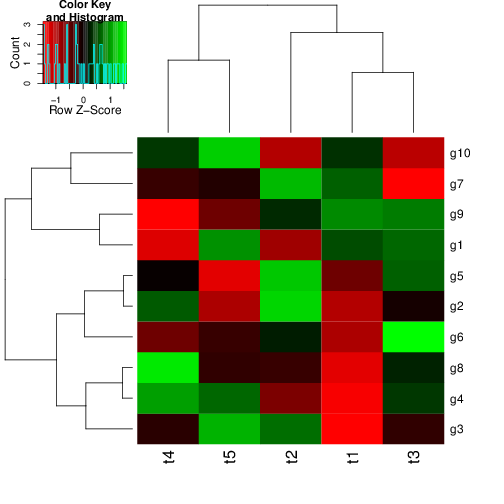
hclust() and agnes()diana()The code sections in this tutorial make use of the following R/Bioc packages: c("ggplot2", "gplots", "pheatmap", "RColorBrewer", "ape", "ComplexHeatmap", "scatterplot3d"). If they are not available on a system, then they need to be installed with BiocManager::install() first.
hclust and heatmap.2Note, with large data sets consider using flashClust which is a fast implementation of hierarchical clustering.
## Row- and column-wise clustering
hr <- hclust(as.dist(1-cor(t(y), method="pearson")), method="complete")
hc <- hclust(as.dist(1-cor(y, method="spearman")), method="complete")
## Tree cutting
mycl <- cutree(hr, h=max(hr$height)/1.5); mycolhc <- rainbow(length(unique(mycl)), start=0.1, end=0.9); mycolhc <- mycolhc[as.vector(mycl)]
## Plot heatmap
mycol <- colorpanel(40, "darkblue", "yellow", "white") # or try redgreen(75)
heatmap.2(y, Rowv=as.dendrogram(hr), Colv=as.dendrogram(hc), col=mycol, scale="row", density.info="none", trace="none", RowSideColors=mycolhc) 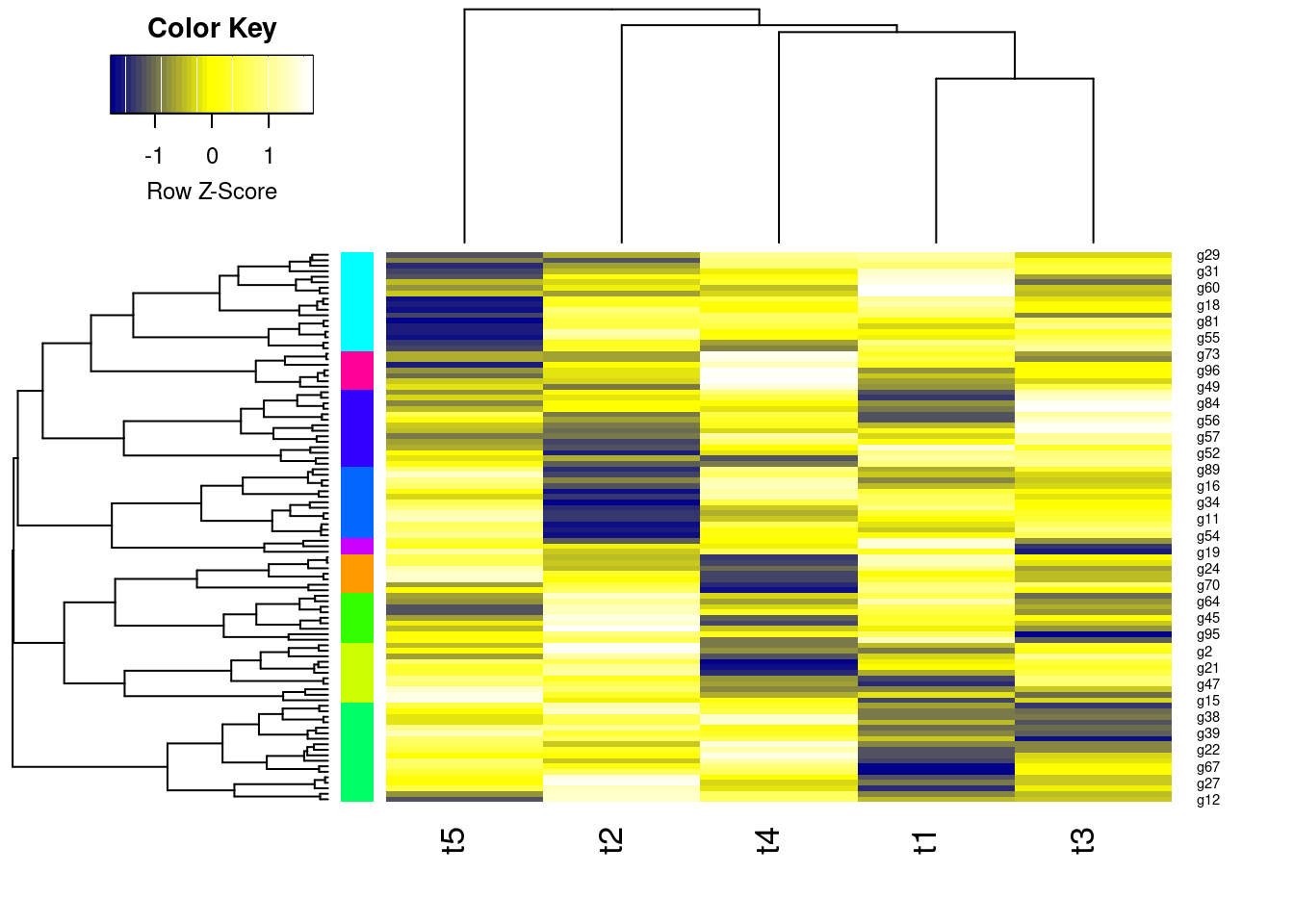
g1 g2 g3 g4 g5 g6 g7 g8 g9 g10 g11 g12 g13 g14 g15 g16 g17 g18 g19 g20 g21 g22 g23 g24 g25 g26 g27 g28 g29 g30 g31 g32 g33 g34 g35 g36 g37 g38 g39 g40 g41 g42 g43 g44
2 3 1 1 1 1 1 2 1 2 2 3 1 2 1 3 3 2 2 1 1 3 3 2 2 1 3 1 1 3 3 1 3 1 3 3 1 2 2 2 2 1 3 3
g45 g46 g47 g48 g49 g50 g51 g52 g53 g54 g55 g56 g57 g58 g59 g60 g61 g62 g63 g64 g65 g66 g67 g68 g69 g70 g71 g72 g73 g74 g75 g76 g77 g78 g79 g80 g81 g82 g83 g84 g85 g86 g87 g88
2 2 1 1 1 1 2 2 3 3 2 2 3 2 2 1 1 1 3 2 2 3 1 1 2 2 2 3 2 1 3 3 2 3 2 1 1 3 3 3 1 1 2 3
g89 g90 g91 g92 g93 g94 g95 g96 g97 g98 g99 g100
2 1 1 2 3 1 1 1 3 2 2 2 fannylibrary(cluster) # Loads the cluster library.
fannyy <- fanny(y, k=4, metric = "euclidean", memb.exp = 1.2)
round(fannyy$membership, 2)[1:4,] [,1] [,2] [,3] [,4]
g1 0.74 0.05 0.07 0.15
g2 0.03 0.84 0.05 0.08
g3 0.23 0.05 0.59 0.13
g4 0.01 0.01 0.93 0.05 g1 g2 g3 g4 g5 g6 g7 g8 g9 g10 g11 g12 g13 g14 g15 g16 g17 g18 g19 g20 g21 g22 g23 g24 g25 g26 g27 g28 g29 g30 g31 g32 g33 g34 g35 g36 g37 g38 g39 g40 g41 g42 g43 g44
1 2 3 3 3 4 4 1 4 4 4 2 3 1 4 4 1 1 4 4 4 1 3 1 1 4 1 3 3 4 1 1 2 4 2 2 3 1 4 1 1 3 1 2
g45 g46 g47 g48 g49 g50 g51 g52 g53 g54 g55 g56 g57 g58 g59 g60 g61 g62 g63 g64 g65 g66 g67 g68 g69 g70 g71 g72 g73 g74 g75 g76 g77 g78 g79 g80 g81 g82 g83 g84 g85 g86 g87 g88
1 1 3 3 4 4 1 4 2 1 1 1 2 4 4 4 3 4 2 1 4 1 3 3 1 1 1 2 4 4 2 2 4 2 4 4 3 2 2 2 3 4 4 2
g89 g90 g91 g92 g93 g94 g95 g96 g97 g98 g99 g100
1 3 3 4 2 3 3 3 2 1 1 1 [,1] [,2] [,3] [,4]
g1 TRUE FALSE FALSE FALSE
g2 FALSE TRUE FALSE FALSE
g3 TRUE FALSE TRUE FALSE
g4 FALSE FALSE TRUE FALSE$g1
[1] 1
$g2
[1] 2
$g3
[1] 1 3
$g4
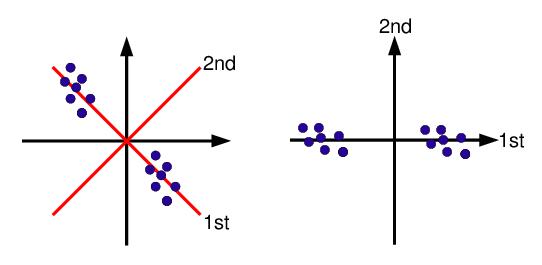
[1] 3Principal components analysis (PCA) is a data reduction technique that allows to simplify multidimensional data sets to 2 or 3 dimensions for plotting purposes and visual variance analysis.
Importance of components:
PC1 PC2 PC3 PC4 PC5
Standard deviation 1.1715 1.0232 0.9814 0.9439 0.8524
Proportion of Variance 0.2745 0.2094 0.1926 0.1782 0.1453
Cumulative Proportion 0.2745 0.4839 0.6765 0.8547 1.0000plot(pca$x, pch=20, col="blue", type="n") # To plot dots, drop type="n"
text(pca$x, rownames(pca$x), cex=0.8)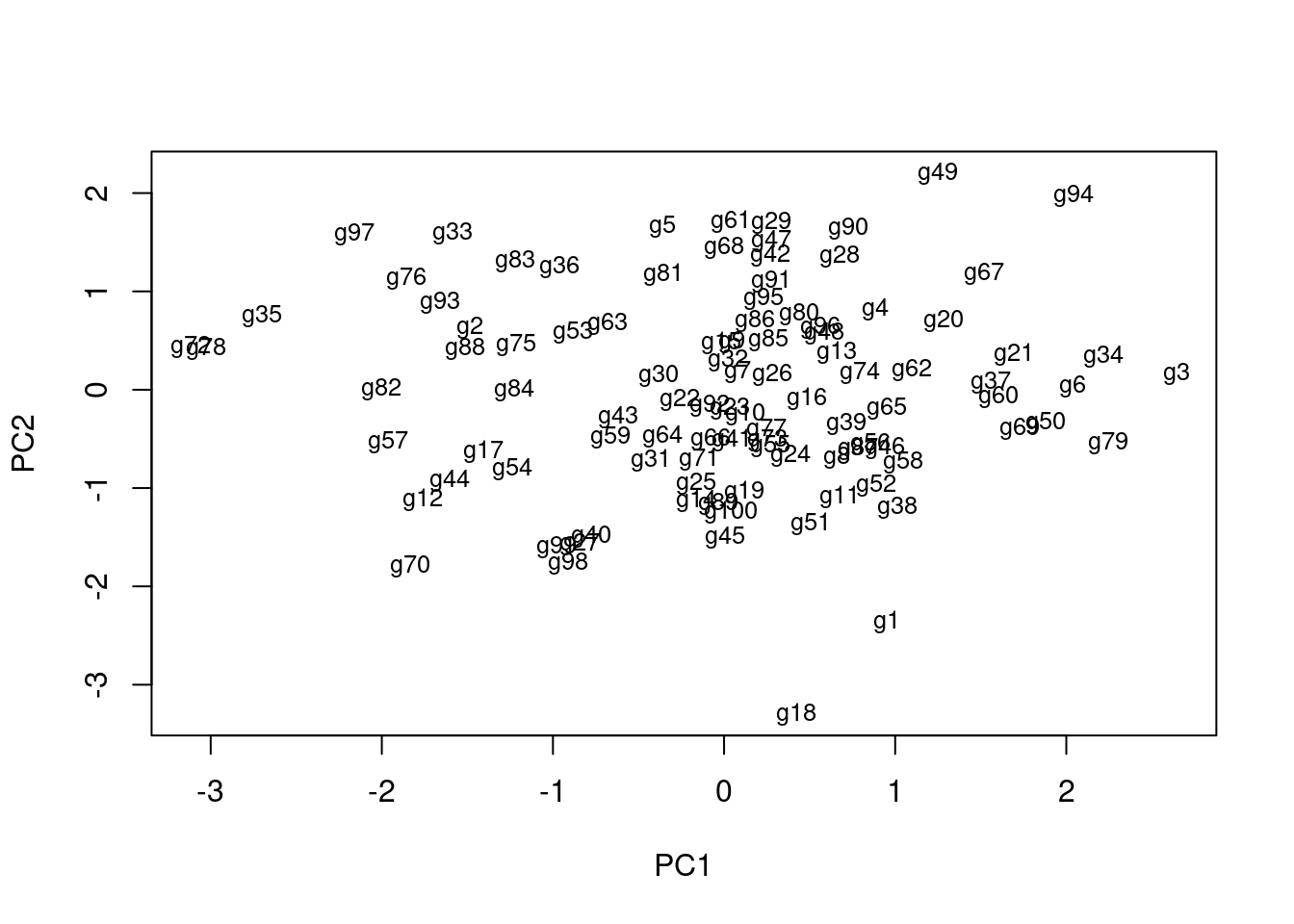
1st and 2nd principal components explain x% of variance in data.
The following example performs MDS analysis with cmdscale on the geographic distances among European cities.
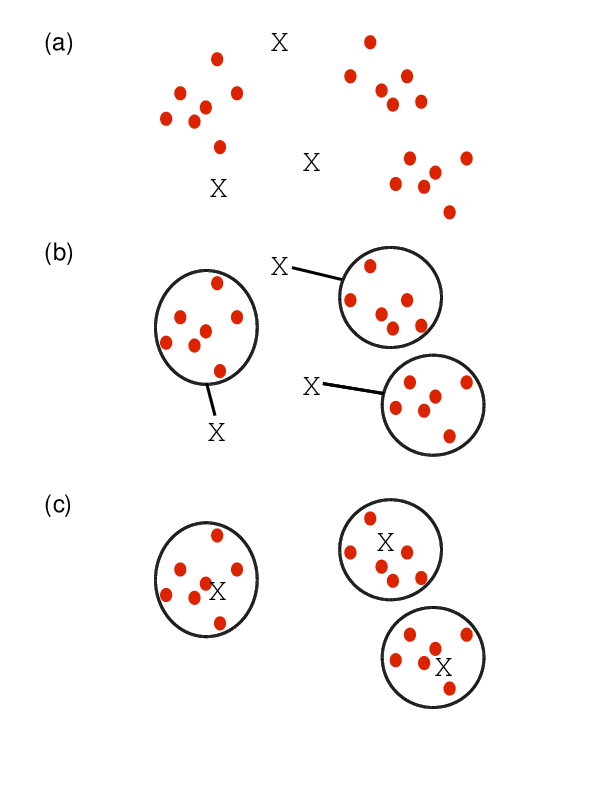
Finds in matrix subgroups of rows and columns which are as similar as possible to each other and as different as possible to the remaining data points.
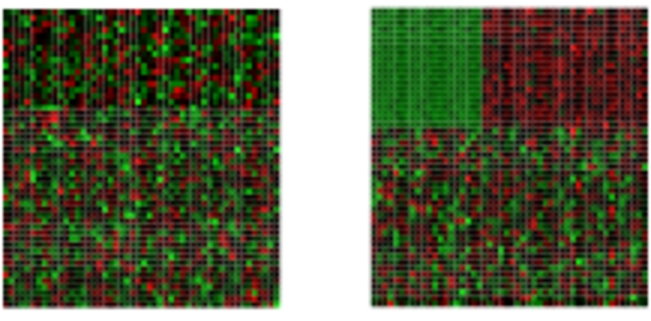
Graph-based clustering methods such as Louvain and Leiden, together with UMAP visualization, have become the standard approach for identifying cell type populations in single-cell RNA-Seq (scRNA-Seq) data. Because these methods are tightly integrated with embedding and dimensionality reduction steps specific to the scRNA-Seq workflow, they are covered in the dedicated scRNA-Seq tutorial:
scRNA-Seq Embedding Algorithm and Clustering Methods
That tutorial covers:
The following imports the cindex() function and computes the Jaccard Index for two sample cluster results.
source("http://faculty.ucr.edu/~tgirke/Documents/R_BioCond/My_R_Scripts/clusterIndex.R")
library(cluster); y <- matrix(rnorm(5000), 1000, 5, dimnames=list(paste("g", 1:1000, sep=""), paste("t", 1:5, sep=""))); clarax <- clara(y, 49); clV1 <- clarax$clustering; clarax <- clara(y, 50); clV2 <- clarax$clustering
ci <- cindex(clV1=clV1, clV2=clV2, self=FALSE, minSZ=1, method="jaccard")
ci[2:3] # Returns Jaccard index and variables used to compute it $variables
a b c
11855 873 558
$Jaccard_Index
[1] 0.8922926The following example shows how one can cluster entire cluster result sets. First, 10 sample cluster results are created with Clara using k-values from 3 to 12. The results are stored as named clustering vectors in a list object. Then a nested sapply loop is used to generate a similarity matrix of Jaccard Indices for the clustering results. After converting the result into a distance matrix, hierarchical clustering is performed with hclust.}
clVlist <- lapply(3:12, function(x) clara(y[1:30, ], k=x)$clustering); names(clVlist) <- paste("k", "=", 3:12)
d <- sapply(names(clVlist), function(x) sapply(names(clVlist), function(y) cindex(clV1=clVlist[[y]], clV2=clVlist[[x]], method="jaccard")[[3]]))
hv <- hclust(as.dist(1-d))
plot(as.dendrogram(hv), edgePar=list(col=3, lwd=4), horiz=T, main="Similarities of 10 Clara Clustering Results for k: 3-12") 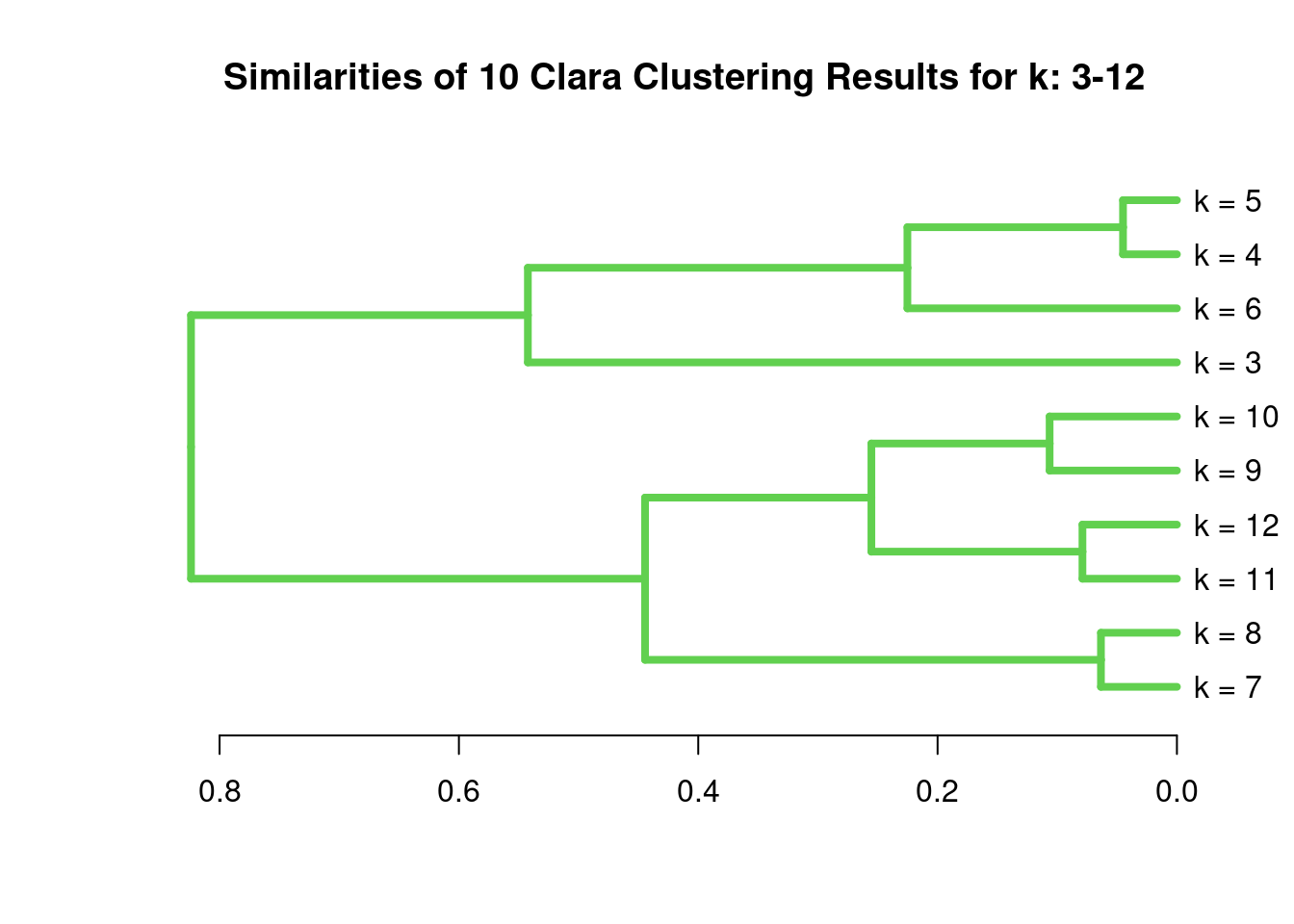
Correlation matrix
g1 g2 g3 g4
g1 1.00000000 -0.2965885 -0.00206139 -0.4042011
g2 -0.29658847 1.0000000 -0.91661118 -0.4512912
g3 -0.00206139 -0.9166112 1.00000000 0.7435892
g4 -0.40420112 -0.4512912 0.74358925 1.0000000Correlation-based distance matrix
hclustHierarchical clustering with complete linkage and basic tree plotting
[1] "merge" "height" "order" "labels" "method" "call" "dist.method"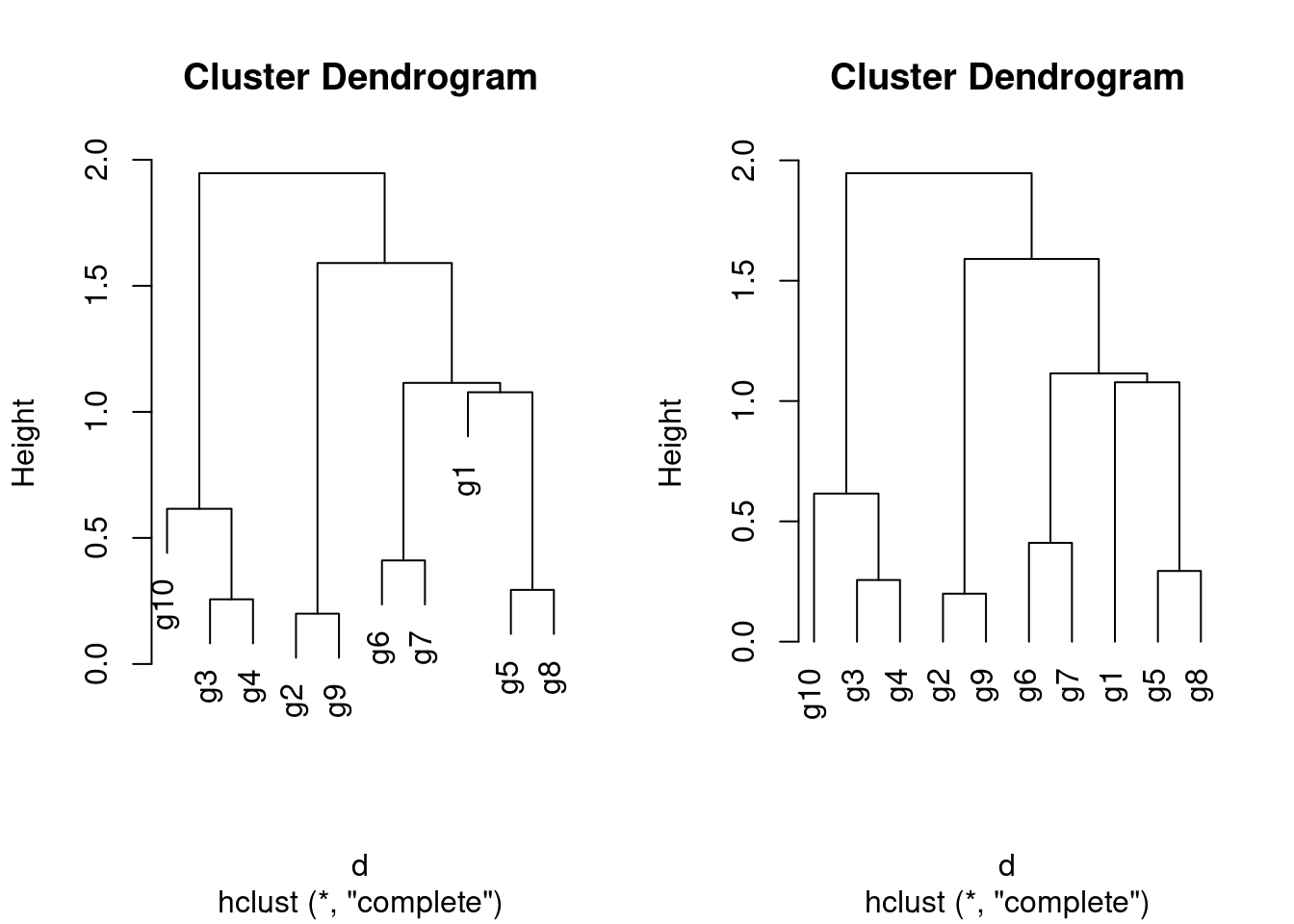
The ape library provides more advanced features for tree plotting
Accessing information in hclust objects
Call:
hclust(d = d, method = "complete", members = NULL)
Cluster method : complete
Number of objects: 10 [1] "g10" "g3" "g4" "g2" "g9" "g6" "g7" "g1" "g5" "g8" Tree cutting with cutree
heatmap.2All in one step: clustering and heatmap plotting
pheatmapAll in one step: clustering and heatmap plotting
Customizes row and column clustering and shows tree cutting result in row color bar. Additional color schemes can be found here.
For plotting complex heatmaps, the ComplexHeatmap provides useful functionalities. The following creates a sample heatmap. For additional examples, visit the manual here.
library(ComplexHeatmap)
set.seed(123)
mat = matrix(rnorm(100), 10)
rownames(mat) = paste0("R", 1:10)
colnames(mat) = paste0("C", 1:10)
column_ha = HeatmapAnnotation(foo1 = runif(10), bar1 = anno_barplot(runif(10)))
row_ha = rowAnnotation(foo2 = runif(10), bar2 = anno_barplot(runif(10)))
Heatmap(mat, name = "mat", top_annotation = column_ha, right_annotation = row_ha)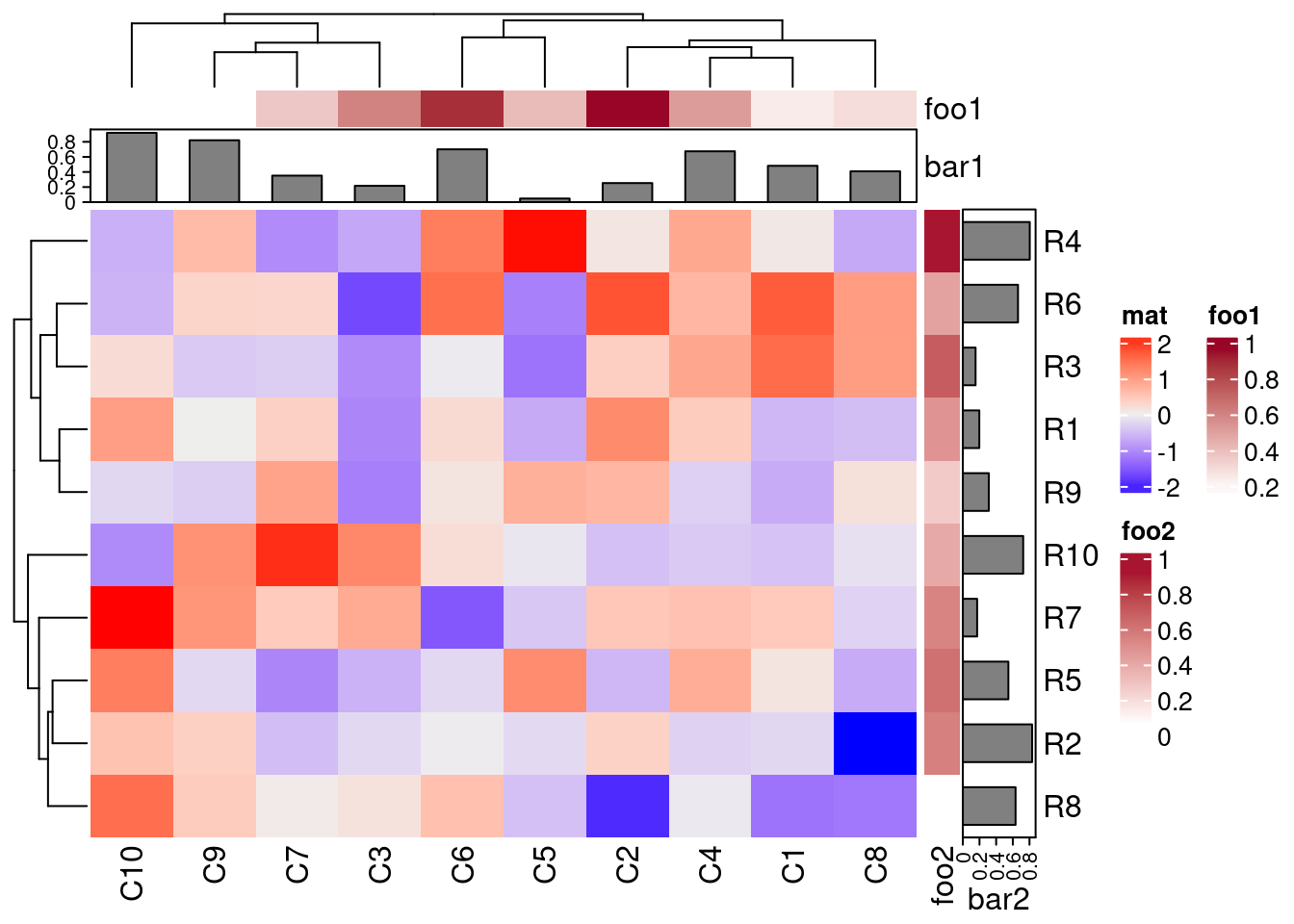
Runs K-means clustering with PAM (partitioning around medoids) algorithm and shows result in color bar of hierarchical clustering result from before.
Performs k-means fuzzy clustering
[,1] [,2] [,3] [,4]
g1 1.00 0.00 0.00 0.00
g2 0.00 0.99 0.00 0.00
g3 0.02 0.01 0.95 0.03
g4 0.00 0.00 0.99 0.01 g1 g2 g3 g4 g5 g6 g7 g8 g9 g10
1 2 3 3 4 4 4 4 2 3 ## Returns multiple cluster memberships for coefficient above a certain
## value (here >0.1)
fannyyMA <- round(fannyy$membership, 2) > 0.10
apply(fannyyMA, 1, function(x) paste(which(x), collapse="_")) g1 g2 g3 g4 g5 g6 g7 g8 g9 g10
"1" "2" "3" "3" "4" "4" "4" "2_4" "2" "3" Performs MDS analysis on the geographic distances between European cities
Performs PCA analysis after scaling the data. It returns a list with class prcomp that contains five components: (1) the standard deviations (sdev) of the principal components, (2) the matrix of eigenvectors (rotation), (3) the principal component data (x), (4) the centering (center) and (5) scaling (scale) used.
[1] "sdev" "rotation" "center" "scale" "x" Importance of components:
PC1 PC2 PC3 PC4 PC5
Standard deviation 1.3611 1.1777 1.0420 0.69264 0.4416
Proportion of Variance 0.3705 0.2774 0.2172 0.09595 0.0390
Cumulative Proportion 0.3705 0.6479 0.8650 0.96100 1.0000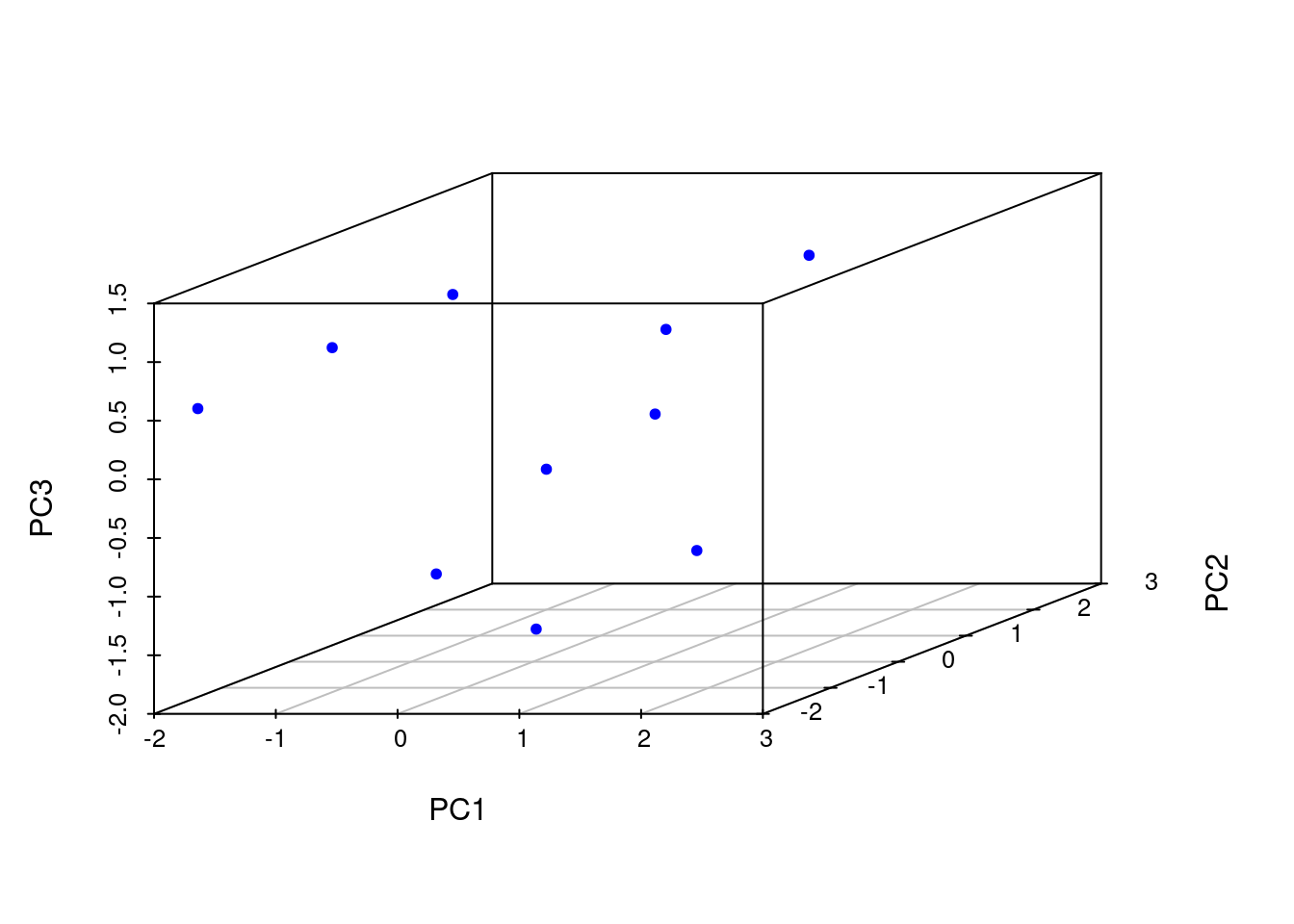
See here
R version 4.5.3 (2026-03-11)
Platform: x86_64-pc-linux-gnu
Running under: Debian GNU/Linux 12 (bookworm)
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.11.0
LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.11.0 LAPACK version 3.11.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8 LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8 LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: America/Los_Angeles
tzcode source: system (glibc)
attached base packages:
[1] grid stats graphics grDevices utils datasets methods base
other attached packages:
[1] scatterplot3d_0.3-44 ComplexHeatmap_2.26.1 RColorBrewer_1.1-3 pheatmap_1.0.13 cluster_2.1.8.2 gplots_3.3.0 ape_5.8-1 ggplot2_4.0.2
loaded via a namespace (and not attached):
[1] generics_0.1.4 bitops_1.0-9 shape_1.4.6.1 KernSmooth_2.23-26 gtools_3.9.5 lattice_0.22-9 digest_0.6.39 magrittr_2.0.4 caTools_1.18.3 evaluate_1.0.5
[11] iterators_1.0.14 circlize_0.4.17 fastmap_1.2.0 foreach_1.5.2 doParallel_1.0.17 jsonlite_2.0.0 GlobalOptions_0.1.3 scales_1.4.0 codetools_0.2-20 cli_3.6.5
[21] rlang_1.1.7 crayon_1.5.3 withr_3.0.2 yaml_2.3.12 otel_0.2.0 tools_4.5.3 parallel_4.5.3 dplyr_1.2.0 colorspace_2.1-2 BiocGenerics_0.56.0
[31] GetoptLong_1.1.0 vctrs_0.7.1 R6_2.6.1 png_0.1-8 magick_2.9.0 stats4_4.5.3 matrixStats_1.5.0 lifecycle_1.0.5 S4Vectors_0.48.0 IRanges_2.44.0
[41] htmlwidgets_1.6.4 clue_0.3-68 pkgconfig_2.0.3 pillar_1.11.1 gtable_0.3.6 glue_1.8.0 Rcpp_1.1.1 xfun_0.56 tibble_3.3.1 tidyselect_1.2.1
[51] knitr_1.51 dichromat_2.0-0.1 rjson_0.2.23 farver_2.1.2 htmltools_0.5.9 nlme_3.1-168 rmarkdown_2.30 Cairo_1.7-0 compiler_4.5.3 S7_0.2.1