Projects
The Girke lab develops computational methods and software at the interface of genome biology, chemical genomics, and bioinformatics. Below is a brief overview of selected research projects, with each project highlighting a representative theme of the lab’s work.
systemPipeR: Reproducible Workflow Management for Data-Intensive Life Science Research
Modern life science research increasingly relies on complex, multi-step computational analyses of large and heterogeneous datasets, creating major challenges for workflow organization, reproducibility, and transparency. systemPipeR is a workflow management environment that enables researchers to design, execute, monitor, and document sophisticated data analysis pipelines within a unified framework. It integrates statistical analysis in R with widely used bioinformatics software and supports execution on personal computers as well as high-performance computing environments. By combining workflow visualization, scalable execution, and automated analysis reporting, systemPipeR helps make modern data-driven research more efficient, reproducible, and standardized.
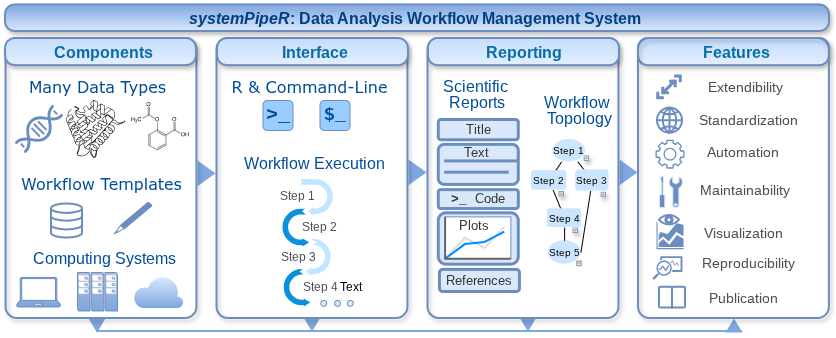
Visual overview of the systemPipeR workflow framework.
Interventions Promoting Healthy Aging and Longevity
The Girke lab develops AI-based strategies to identify drugs, natural compounds, and combination therapies that promote healthy aging and delay age-related disease. By integrating large-scale omics signatures, genetic and pharmacological perturbation data, and new methods for drug–target discovery, this work seeks to uncover more effective and sustainable interventions for extending health span.
Experimental validation is performed in collaboration with consortium scientists using mouse models and human iPSC systems, including groups at the Salk Institute and the University of Michigan. These combined efforts aim to translate computational discoveries into more precise and personalized approaches for healthier aging in humans. This research is supported by a cooperative U19 grant from the National Institute on Aging at the National Institutes of Health (U19AG023122).
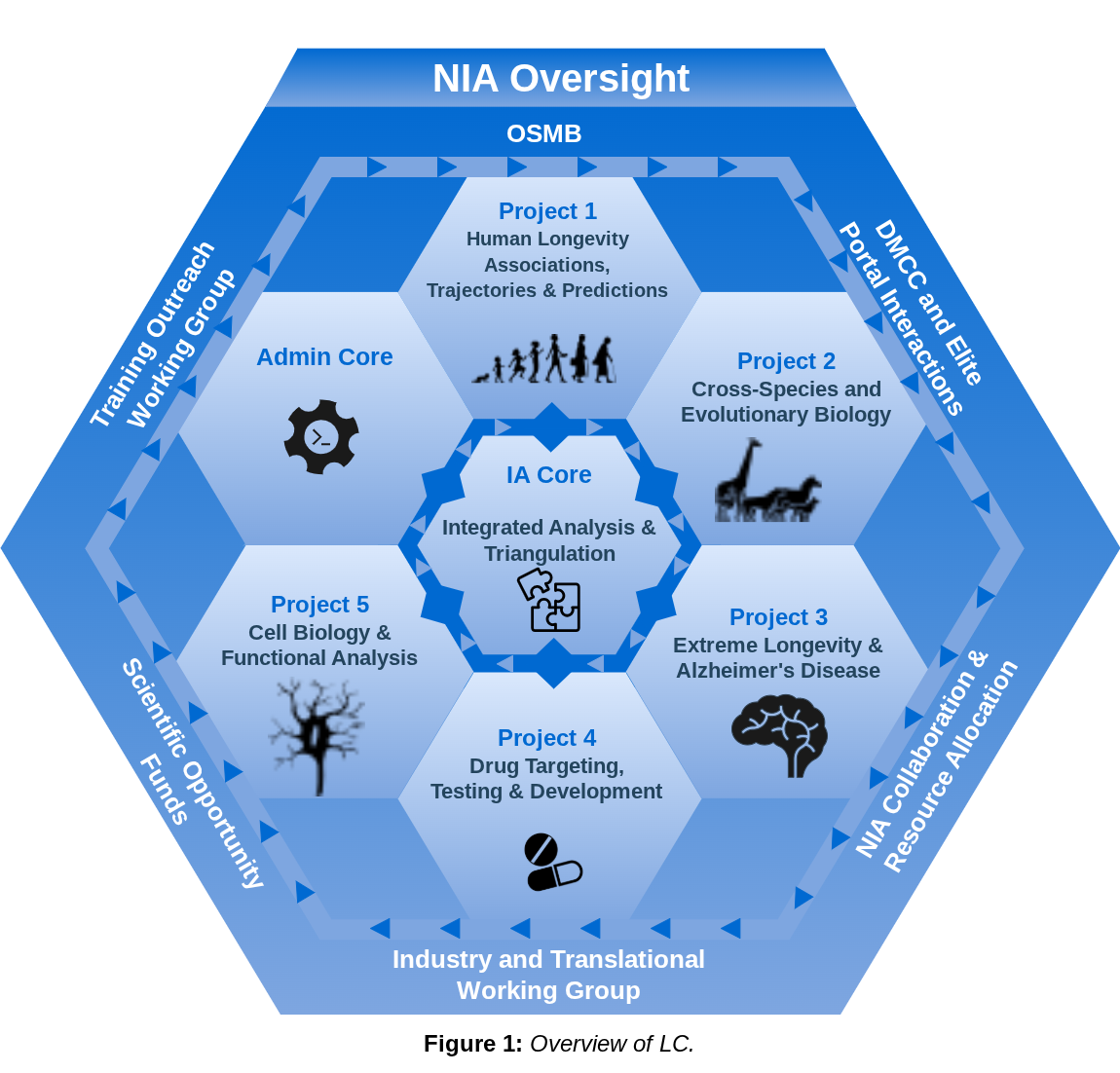
Overview of Longevity Consortium.
spatialHeatmap: Visualizing Spatial Bulk and Single-Cell Assays in Anatomical Images
The spatialHeatmap package enables intuitive visualization of single-cell, spatial, and other tissue-resolved omics data by overlaying quantitative assay values onto anatomical images. It combines spatial heatmaps with complementary clustering and network-based views, and is available both as an R package and as an interactive Shiny application for exploratory analysis by computational and experimental users alike.
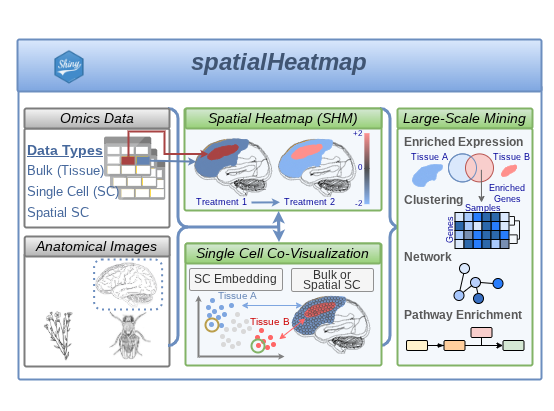
Spatial data visualization in anatomical images.
Gene Expression Searching with signatureSearch
The signatureSearch package provides an integrated R/Bioconductor environment for searching gene expression signatures across large reference databases derived from disease states, genetic backgrounds, chemical perturbations, and single-cell datasets. By combining signature matching with functional enrichment analysis and network-based visualization, it helps reveal related cellular responses, identify perturbed biological processes, and support discovery of novel drug, pathway, and cell-state connections.
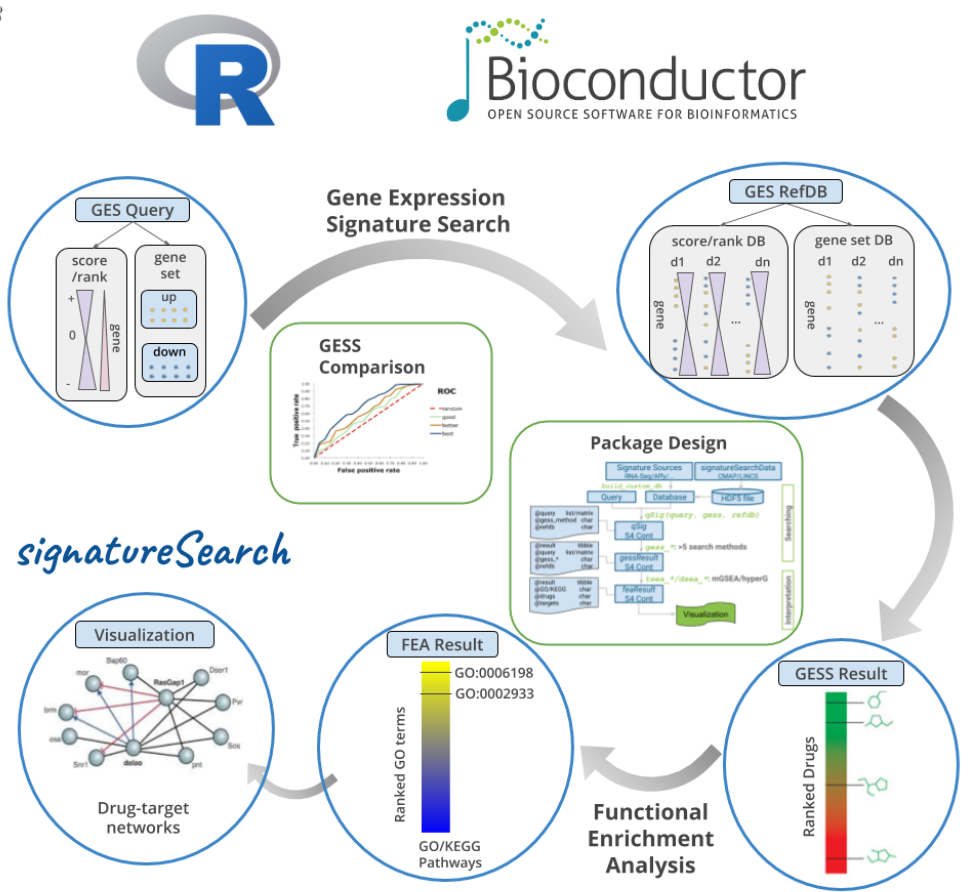
Signature searching.
Software for Small Molecule Discovery and Chemical Genomics
The ChemmineR environment is a modular software infrastructure designed for modeling similarities among drug-like small molecules and high-throughput screening data in drug discovery and chemical genomics. It consists of five R/Bioconductor packages and the user-friendly ChemMine Tools web interface, providing utilities for molecule processing, property prediction, structural similarity searching, and clustering. A key component is the fmcsR algorithm, which provides mismatch-tolerant maximum common substructure searches and has demonstrated superior virtual screening performance. Related software and resources include ChemmineOB and eiR.
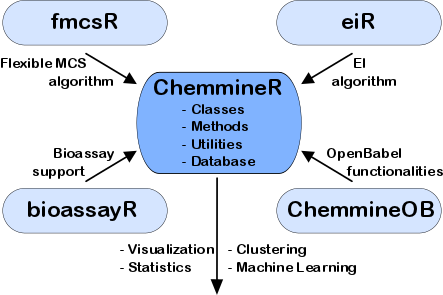
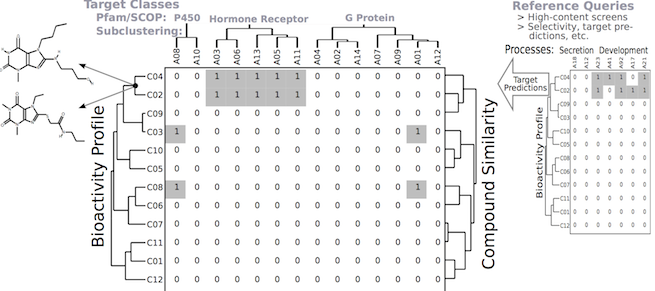
Chemical genomics.
Drug-target interaction analysis
The drugTargetInteractions package provides tools for identifying known and candidate interactions between small molecules and their gene or protein targets. By leveraging the extensive bioactivity information available in the public domain, including the ChEMBL database, it supports efficient mapping of drug–target relationships for applications in chemical biology, target discovery, and translational drug research.
Software: Bioc
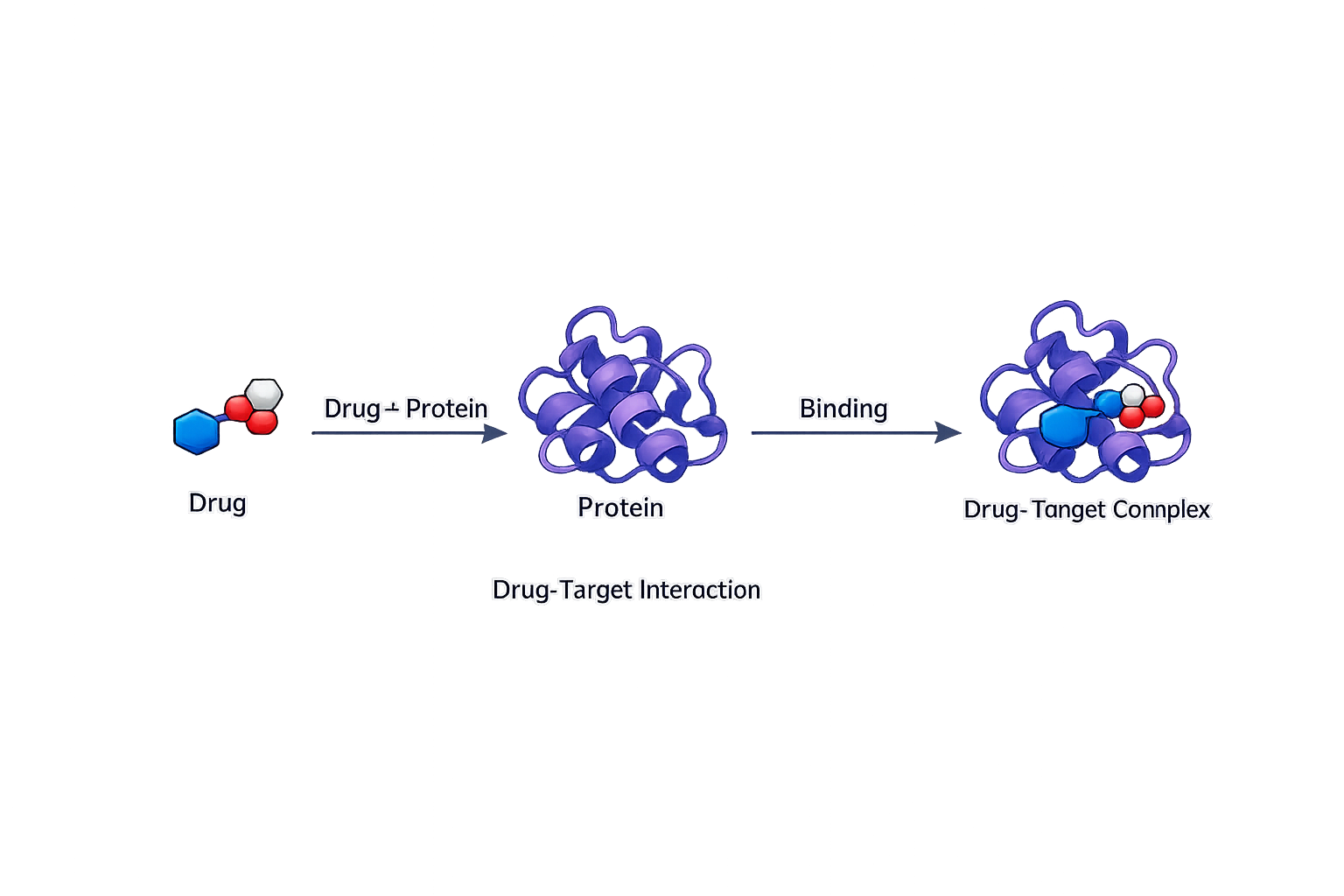
Drug-target interaction mining.
Large-scale bioactivity analysis
This project used large-scale bioactivity data to compare how approved drugs and other small molecules interact with many different protein targets. The study uncovered numerous promising new drug and target candidates while also helping distinguish compounds with selective activities from those with broad, less specific effects, providing a valuable resource for future drug discovery efforts.
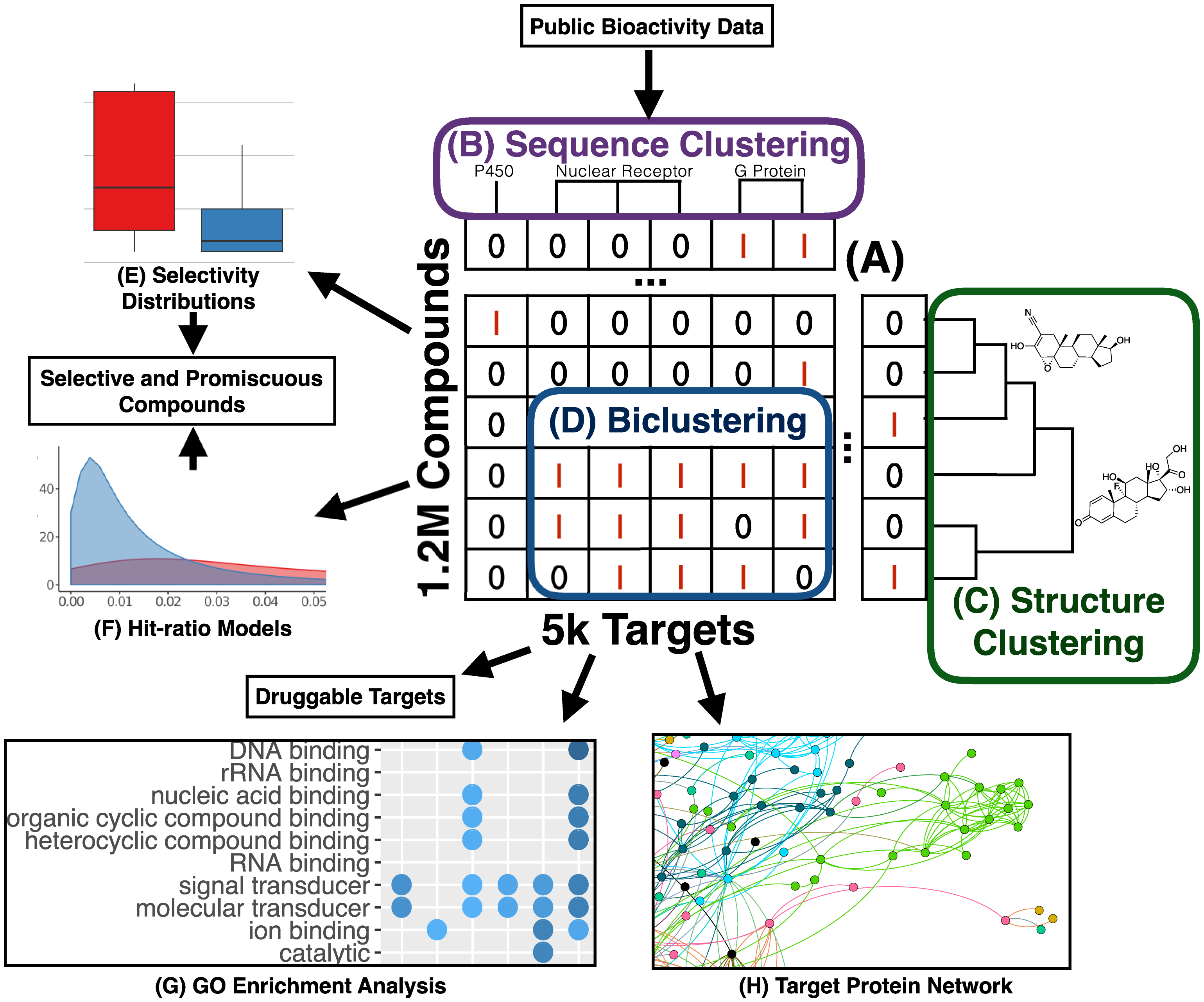
Bioactivity data mining strategy.